|
A Database of Potassium Ion Channel Homology Models & Molecular Dynamics Simulations
Philip W. Fowler, Anthony Ivetac, Syma Khalid, Younes Mokrab, Phillip J. Stansfeld, Kaihsu Tai & Mark S.P. Sansom
Many biophysical studies indicate interactions with lipid/detergent molecules
are critical to the folding/stability of membrane proteins, but only limited
structural data are available on these interactions.
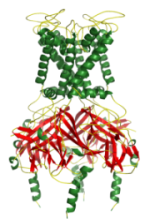
 Data from MD simulations of K channels (both X ray structures and homology models), has been generated, collected and analysed using the following protocol:
Data from MD simulations of K channels (both X ray structures and homology models), has been generated, collected and analysed using the following protocol:
» Embed the K channel (X-ray structure or homology model) in a phospholipid bilayer, solvate and add ions
» Carry out minimum of 10 ns of MD using GROMACS
» Extract snapshot of each protein in its optimum position within bilayer
» Submit to database: simulation properties/snapshots, final system coordinates, and analysis of lipid-protein interactions
 This work has been supported by:
This work has been supported by:
» BioSimGrid (molecular
dynamics trajectory database)
» Biotechnology and Biological
Sciences Research Council, Wellcome Trust (funding)
» Alessandro Grotessi, Zara Sands, Shozeb Haider (simulations)
|